The Philips raw data reader node, for MR data, is now available as a binary release. Users of the Philips raw data formats can directly import the raw data into GPI and start investigating. Thanks to the tenacious efforts of Ryan Robison, the reader node supports a plethora of file formats such as .data, .list, .lab, .raw, .sin, .par, .xml, .rec, and .cpx at many different release levels. The package releases are available for download on GitHub.
Output
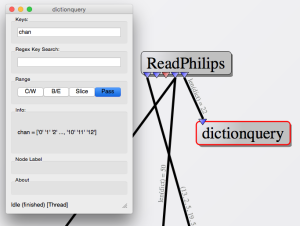
The ReadPhilips node parses the file contents for MR data, converts the data to a numpy numeric array and makes it available as an output port. Depending on the input files and application, there may be multiple output datasets corresponding to the sampled k-space, noise measurements, etc... The reader also parses any available header information (which is format dependent) and populates a python-dictionary object which is also pushed to an output port. This header information can be viewed via the 'dictionquery' node which displays python-dictionary information as a simple list.
Widgets
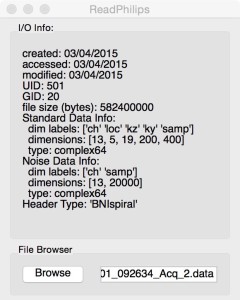
The ReadPhilips menu provides an interface to browse for the desired files and displays basic file system information and a summary of the header information (when available). The interface also provides options for various minor data corrections and the ability to load single coils or slices from the file.
Community
The ReadPhilips node is available to the scientific community as a closed source binary package allowing users immediate access to this functionality. The source code for this project can be made available to Philips researchers, who have signed an NDA, in an effort to foster community development and maintenance of the reader as the Philips product software advances. This project provides a model for GPI developers to generate reader tools for other closed raw data formats. In this way, the closed source tools can be maintained by developers, who have vendor specific access, allowing the tool to be updated with product changes. The end users benefit by moving forward with their research without getting tangled up in the proprietary code.
If you have a GPI node project, you are welcome to submit a link for listing on the community page.